 |
The big picture
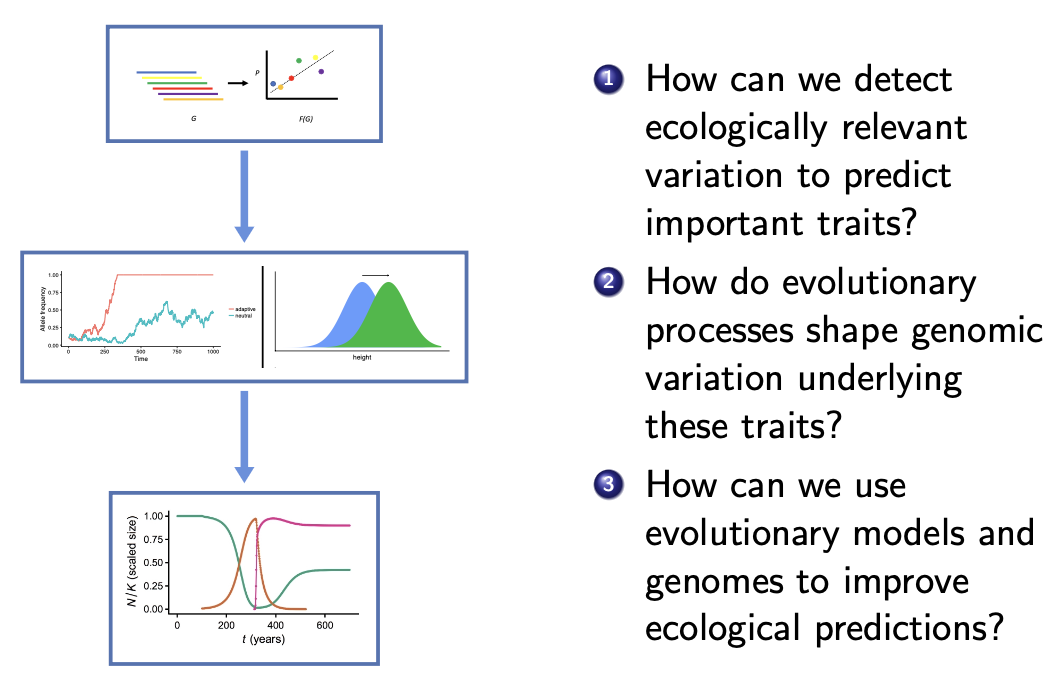
Ecology has developed a powerful set of analytical tools that can characterize how changes in the environment affect species'
abundance,
range, and community composition.
Population genetics has developed a similarly powerful set of tools for relating genomic
patterns to past evolutionary events.
Our lab seeks to combine the forward-looking perspective of population/community ecology
with mechanistic models of evolution to characterize how species evolve in response to environmental change. We integrate
mathematical models with experimental and genomic data in pursuit of this goal.
Detecting adaptation in response to changing environments
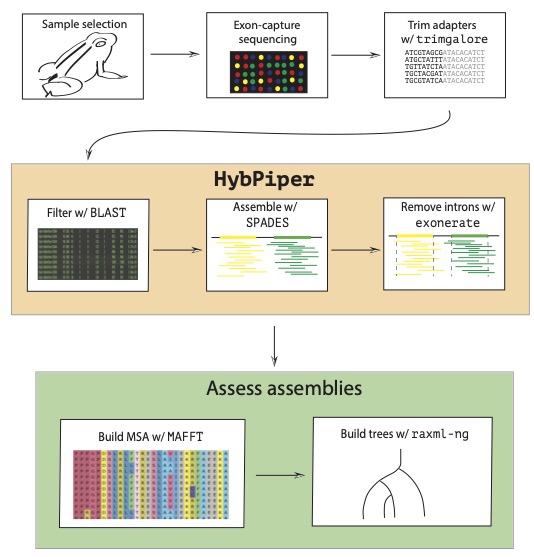 Rare alleles can have very large effects on individual phenotypes and fitness, but are typically hard to detect in association studies.
Modern genomic data show that alleles can have small or large effects on both traits and fitness. The effect sizes and frequency of these alleles matters, because
species may be able to evolve much faster than anticipated in response to environmental change if large effect alleles contribute to rapid adaptation.
Our work leverages the power of genomes to develop methods to detect rare allele contributions to local adaptation, detect loci involved in complex traits, and
infer adaptation rates when fitness effects are variable in magnitude. We are especially interested in linking modern genomic data with trait associations. We
use thermal performance of atlantic slipper snails as a model system to study these questions, while simultaneously developing methods and models that are
not species-specific.
Rare alleles can have very large effects on individual phenotypes and fitness, but are typically hard to detect in association studies.
Modern genomic data show that alleles can have small or large effects on both traits and fitness. The effect sizes and frequency of these alleles matters, because
species may be able to evolve much faster than anticipated in response to environmental change if large effect alleles contribute to rapid adaptation.
Our work leverages the power of genomes to develop methods to detect rare allele contributions to local adaptation, detect loci involved in complex traits, and
infer adaptation rates when fitness effects are variable in magnitude. We are especially interested in linking modern genomic data with trait associations. We
use thermal performance of atlantic slipper snails as a model system to study these questions, while simultaneously developing methods and models that are
not species-specific.
Characterizing assembly processes in ecological communities
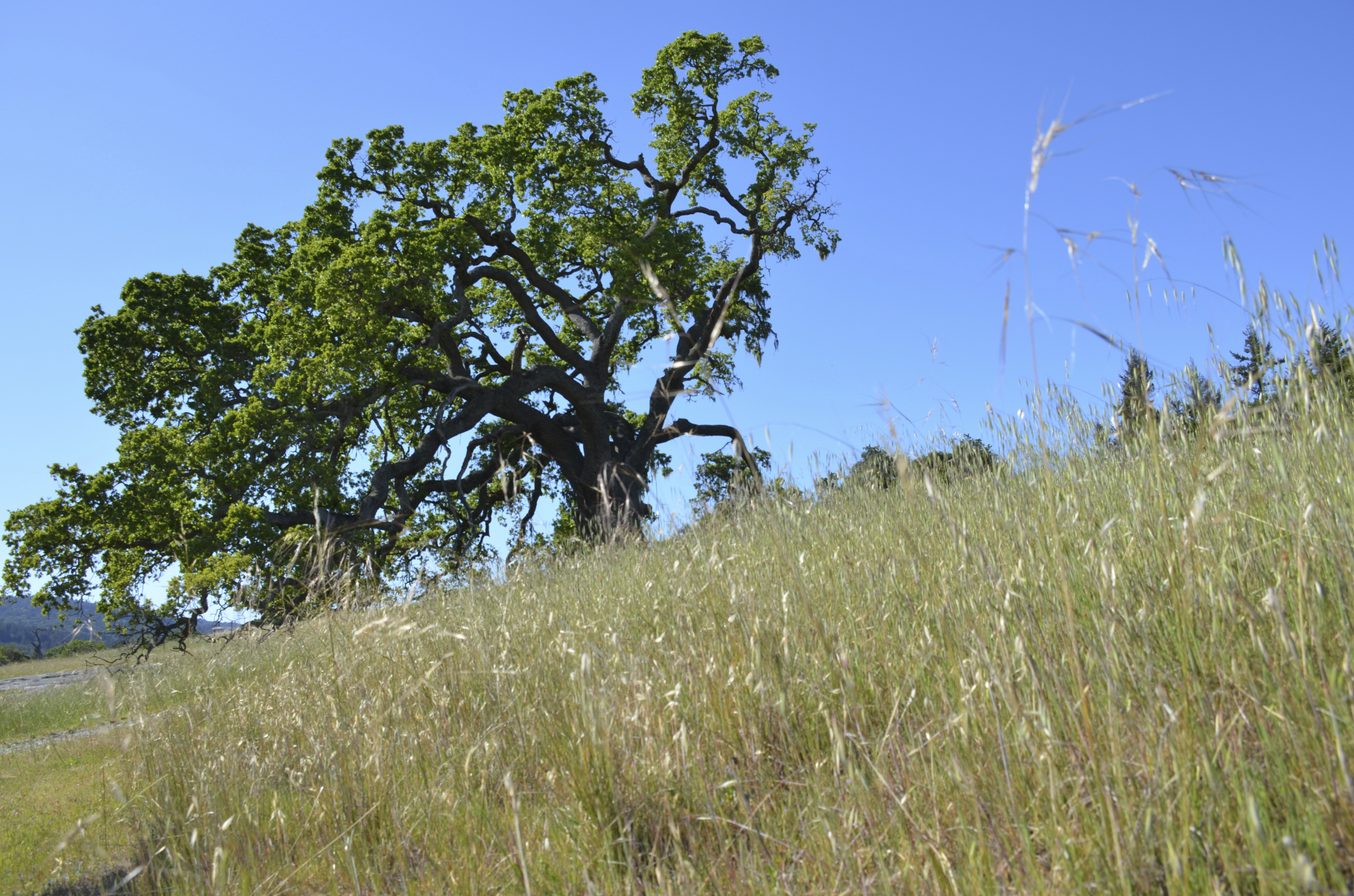 Exotic species can dramatically affect species composition by directly out-competing native species, but in some cases native and introduced species may coexist stably.
We studied competition between native and exotic species in California grasslands. Our analysis suggests
that exotic annual species are liekly to outcompete California native species, but that their likelihood of persistence depends integrally on the order of arrival of their exotic competitors.
Our results provide a contrast to some earlier studies that focused on undisturbed communities and found stabilizing niche differences between species, perhaps due to the long co-evolutionary history of naturally co-occurring species. Studies like this one can help inform our thinking about the processes that
could explain diversity in nature, but we still know so little about how biotic interactions affect species' distributions and persistence!
Exotic species can dramatically affect species composition by directly out-competing native species, but in some cases native and introduced species may coexist stably.
We studied competition between native and exotic species in California grasslands. Our analysis suggests
that exotic annual species are liekly to outcompete California native species, but that their likelihood of persistence depends integrally on the order of arrival of their exotic competitors.
Our results provide a contrast to some earlier studies that focused on undisturbed communities and found stabilizing niche differences between species, perhaps due to the long co-evolutionary history of naturally co-occurring species. Studies like this one can help inform our thinking about the processes that
could explain diversity in nature, but we still know so little about how biotic interactions affect species' distributions and persistence!
Our ongoing work seeks to incorporate genomic data into ecological models to predict when species composition is likely to depend on rapid evolutionary
processes. Evolutionary dynamics could be an imporant component of assembly processes in nature, and could be particularly relevant for species that
experience shifts in their environental context due to climate change or introduction to new habitats. Microbes are an ideal model for asking such questions
because of their short generation times, feasibility of whole-genome sequencing, well-characterized biological interactions, and natural hitory relevance.
Graduate student Kasturi Lele is using a combination of experimental microbial community assembly data, genomic sequecnes, and model-based simulations
to characterize how evolutionary and ecological processes may affect community stability in the sourdough microbiome. She is working in collaboration with
Dr. Ben Wolfe's lab at Tufts.